Saccharum streak virus
Basic Information
| Genus |
Mastrevirus
|
| NCBI Assembly |
GCF_000886895.1 |
| Isolate |
South Africa: KwaZulu-Natal |
| Release date |
2015/2/22 |
| Submitter |
Lawry,R., Martin,D.P., Shepherd,D.N., van Antwerpen,T., Varsani,A. |
| Host |
|
| Vector |
|
| Download |
Genome
|GFF3
|PEP
|CDS |
Genomic Organization
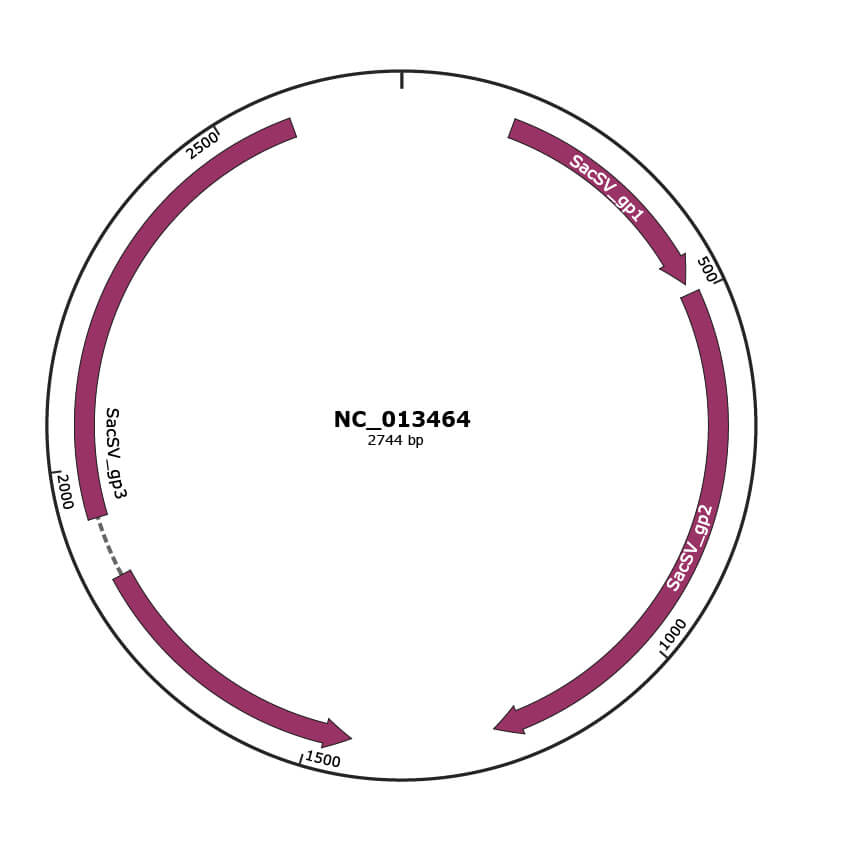
JBrowse
Genome
TAATATTACCGCATCCCCTTTTGCGAGGGCCCCAAAGGCCCGAGCGGTCCTAGCCGCTTTGTCTCTGTGGGGTATTTCGAAGATGAAATCTGCTCTTTCGACCGCCGACAGGCTAAGGTGCGATATAAATCAGCCGTCCTCGCCTTGCTTTGAACATGGAGGGCGCCTACGGTGCGATCTATCCTTCCGCGCAGAGTGCTTTGCCGCGGGTACCCATTGCAGCTCCGTCCTCTCCGAGCTTGCCGTGGAGTCGCGTCGGTGAGATAGCTATCTTTGTCTTTGTTGCAGTACTAAGCTTTTATCTGCTGTGGGTGTGGGTGCTGAGAGATCTTATCTTCGTTGTGAAGGCTCGCCGAGGGCACTCAACGGAGGAGCTGCGCTTTGGCCCTACCGTTCAAGCCCCTCCCGTCGCGCCAGTGTCTGTTCCTGGTGCGTCCGCTGTCACTGCTAGCTGTCCACCGGAGCCTAGACCTTTCTGTGTCTAGGCGGGCCATCAGCTATGTCTTCTTCCCTTGGCAAGAGGAAGAGGTCGAATGGAGGCGATTGGTCTAAGCGCTCCGCTAAGAAGAAGCCGGCGGGTACCCCTTCACGCCGTGCTGGGCCTGGAAGAGGCCCACGTCCAGCTCTACAGATTGCGACCTACCAGGCCGCTGGAACCTCTATGGTTACTGTCCCTAGCGGGGGCGTTTGTGAACTCCTTGCGACATATGCTCGAGGGTCCGACGAGGGCAACCGTCACACCAACGAGACTATCACGTACAAGGTTGCCTTGGACTACCACTTTGTAGCCTCGTCTGCTGCTTGCAGGTACTCTTCCATCGGTGTGGGGGTCGTGTGGTTGGTGTATGATGCACAGCCCTCCGGCAATGCCCCCCAGGTAACGGATATCTTCCCGCATCCTGATAGTCTGGCCGCGTTCCCGTACACTTGGAAGGTCGGAAGGGAGGTGTGTCATCGCTTCGTGGTGAAGCGGAGGTGGACTTTCACGATGGAGACGGACGGGCGTATCGGGAGCGATATCCCTCGGTCGACGGATTCTTGGCCGCCCTGCAAGCGGTCTATTTACTTTCACAAGTTTGCCACCGGTCTCGGTGTCAAAACGGAGTGGAAGAATCTCGCTGATGGAGGAGTTGGATCCATTAAGAAGGGTGCCCTGTATATTGTAATTGCCCCCGGCAATGGTTTAGAGTTTACGGCGCATGGCAATGCCCGTTTGTACTTTAAGTCTGTTGGAAATCAGTGATTTCCACGAAATAATAAATAAATTTATTTATTACAATGATTGGAATGCGTAGCATTACATTACAGTACATAGTCTGCAGATGTGCAGACCCAAACACACACATACCCAACTCGGCGGTCAGATCGTAGGCGGCTAAGGGGTAGGACTCGAAAAAAACACACGAAAAACATGATATTATTATAATTGCAGCCGCCGGCTTTACGCAGGGGTGAACCACTTCTCCCCGGAGCTCATGATGTAGACTTCGCAGTTTGCCTCCATGTAGTCCCGCTGCGCGGGAGTCATGTCCCGCAGCCAGTCCTCATCTTCGTTGGCGAGGATTACAGCAGGAATGGAGCGTTTAGCGACCTTCTTTTTCTTCCCGTATTTCGGGTTTACAACGTAGTTCTTCTGACAGCCAACGAGCTGCTTCCAGCAAGGACAGTATTTAAAGGGAATGTCATCTATGACATTCAGGACTGCTTCCTCGTCATATGAGGACCAGTCCACGTTATTCTGCCAGTAGTTGTGGCGACCTAGGCTTCTCGCCCAGGTGGATTTGCCGGTCCTTGTTGGACCGACGATGTACAGGCTTCGCGCTCTTCCGGCCTCTTGAGGTTGCTGTGATACAGTTCGTCGAGCCAGACAAGGTCAGAGGTTGCCTGTTCTAAAGAGCAGCCATGAAGAAGAGTATAGTCCTCAGGAGTGACCTGATAGACGTTGAAGTTCAGCCATTCTTCAATGCGCTCGTAGCAGTGGAGGTCGGGTTGTGTTGGTGGATGAGGATTGGTGTAGGGCTCTGCTATCTCAGGGAACAGCTTATTAGCTGAGTACTCGAAGTACTGCAGCTTCGTTGCCCACTCGTACGGTAGCTCCTTCTGAAGCATGGAGAGGTACTCGTGCTTGTTGGTGGAGTGTTGAATGATATCGCGAACGATATCATCCTTTGAGGCGCGAGTGTTAGAGCTCTCACCGATCTGAGGAACGAAGGGCTTCTTCCGTGGAACGAAAGTACCCTTCTCCCACTGACACAATGGGTCCTTGAGTATGTACTCTCTTACCCTGTCTACCGATTTAGCAGACTGGATGTTGGGGTGGTACTCATTTACGTCGAAGAACCGCGGGTCTGAAGTCGTCACGGGGCGGACGCTCTGTGCCAGGGCATGGCAGTGCCATGTCCCATCTTGATGAGCTTCTCTAGCCACCAGGATGTATGCCGGAGTCCATGGAGCTAGCTTGCTCCATAGGCTCAGACCCAAGATCTCGGGATCAAGGCTACACTTGCTGTAGGTTAGGAAGGTGTTTGCGTTCCTGTGCCTGAAGGAACGAGAGCTGTTACTCTCAGTGCTAGTGCTGTTAGCGTAGGCCATCGGACGGCTGTGGTTGTGCTTTGAGATCCGAGGTTTCTCAAAACTCTAGCTAGACTTCGCATCGAAATCCGCCAGCACCCGGGCGGCTTTTATAGCTGTCTTATATGGGCTGGGCCGAAAGGCCGGCCGGCATACAAAAGGGGATGCGAAGA
Gene Information
|
NCBI Accession
|
YP_003288766.1
|
|
Location
|
156-485 |
|
Protein Name
|
movement protein |
|
Coding Region
|
ATGGAGGGCGCCTACGGTGCGATCTATCCTTCCGCGCAGAGTGCTTTGCCGCGGGTACCCATTGCAGCTCCGTCCTCTCCGAGCTTGCCGTGGAGTCGCGTCGGTGAGATAGCTATCTTTGTCTTTGTTGCAGTACTAAGCTTTTATCTGCTGTGGGTGTGGGTGCTGAGAGATCTTATCTTCGTTGTGAAGGCTCGCCGAGGGCACTCAACGGAGGAGCTGCGCTTTGGCCCTACCGTTCAAGCCCCTCCCGTCGCGCCAGTGTCTGTTCCTGGTGCGTCCGCTGTCACTGCTAGCTGTCCACCGGAGCCTAGACCTTTCTGTGTCTAG |
|
Protein Sequence
|
MEGAYGAIYPSAQSALPRVPIAAPSSPSLPWSRVGEIAIFVFVAVLSFYLLWVWVLRDLIFVVKARRGHSTEELRFGPTVQAPPVAPVSVPGASAVTASCPPEPRPFCV |
|
NCBI Accession
|
YP_003288767.1
|
|
Location
|
500-1243 |
|
Protein Name
|
coat protein |
|
Coding Region
|
ATGTCTTCTTCCCTTGGCAAGAGGAAGAGGTCGAATGGAGGCGATTGGTCTAAGCGCTCCGCTAAGAAGAAGCCGGCGGGTACCCCTTCACGCCGTGCTGGGCCTGGAAGAGGCCCACGTCCAGCTCTACAGATTGCGACCTACCAGGCCGCTGGAACCTCTATGGTTACTGTCCCTAGCGGGGGCGTTTGTGAACTCCTTGCGACATATGCTCGAGGGTCCGACGAGGGCAACCGTCACACCAACGAGACTATCACGTACAAGGTTGCCTTGGACTACCACTTTGTAGCCTCGTCTGCTGCTTGCAGGTACTCTTCCATCGGTGTGGGGGTCGTGTGGTTGGTGTATGATGCACAGCCCTCCGGCAATGCCCCCCAGGTAACGGATATCTTCCCGCATCCTGATAGTCTGGCCGCGTTCCCGTACACTTGGAAGGTCGGAAGGGAGGTGTGTCATCGCTTCGTGGTGAAGCGGAGGTGGACTTTCACGATGGAGACGGACGGGCGTATCGGGAGCGATATCCCTCGGTCGACGGATTCTTGGCCGCCCTGCAAGCGGTCTATTTACTTTCACAAGTTTGCCACCGGTCTCGGTGTCAAAACGGAGTGGAAGAATCTCGCTGATGGAGGAGTTGGATCCATTAAGAAGGGTGCCCTGTATATTGTAATTGCCCCCGGCAATGGTTTAGAGTTTACGGCGCATGGCAATGCCCGTTTGTACTTTAAGTCTGTTGGAAATCAGTGA |
|
Protein Sequence
|
MSSSLGKRKRSNGGDWSKRSAKKKPAGTPSRRAGPGRGPRPALQIATYQAAGTSMVTVPSGGVCELLATYARGSDEGNRHTNETITYKVALDYHFVASSAACRYSSIGVGVVWLVYDAQPSGNAPQVTDIFPHPDSLAAFPYTWKVGREVCHRFVVKRRWTFTMETDGRIGSDIPRSTDSWPPCKRSIYFHKFATGLGVKTEWKNLADGGVGSIKKGALYIVIAPGNGLEFTAHGNARLYFKSVGNQ |
|
NCBI Accession
|
YP_003288768.1
|
|
Location
|
1442-1844,1931-2592 |
|
Protein Name
|
RepA |
|
Coding Region
|
ATGGCCTACGCTAACAGCACTAGCACTGAGAGTAACAGCTCTCGTTCCTTCAGGCACAGGAACGCAAACACCTTCCTAACCTACAGCAAGTGTAGCCTTGATCCCGAGATCTTGGGTCTGAGCCTATGGAGCAAGCTAGCTCCATGGACTCCGGCATACATCCTGGTGGCTAGAGAAGCTCATCAAGATGGGACATGGCACTGCCATGCCCTGGCACAGAGCGTCCGCCCCGTGACGACTTCAGACCCGCGGTTCTTCGACGTAAATGAGTACCACCCCAACATCCAGTCTGCTAAATCGGTAGACAGGGTAAGAGAGTACATACTCAAGGACCCATTGTGTCAGTGGGAGAAGGGTACTTTCGTTCCACGGAAGAAGCCCTTCGTTCCTCAGATCGGTGAGAGCTCTAACACTCGCGCCTCAAAGGATGATATCGTTCGCGATATCATTCAACACTCCACCAACAAGCACGAGTACCTCTCCATGCTTCAGAAGGAGCTACCGTACGAGTGGGCAACGAAGCTGCAGTACTTCGAGTACTCAGCTAATAAGCTGTTCCCTGAGATAGCAGAGCCCTACACCAATCCTCATCCACCAACACAACCCGACCTCCACTGCTACGAGCGCATTGAAGAATGGCTGAACTTCAACGTCTATCAGGTGCAACCTCAAGAGGCCGGAAGAGCGCGAAGCCTGTACATCGTCGGTCCAACAAGGACCGGCAAATCCACCTGGGCGAGAAGCCTAGGTCGCCACAACTACTGGCAGAATAACGTGGACTGGTCCTCATATGACGAGGAAGCAGTCCTGAATGTCATAGATGACATTCCCTTTAAATACTGTCCTTGCTGGAAGCAGCTCGTTGGCTGTCAGAAGAACTACGTTGTAAACCCGAAATACGGGAAGAAAAAGAAGGTCGCTAAACGCTCCATTCCTGCTGTAATCCTCGCCAACGAAGATGAGGACTGGCTGCGGGACATGACTCCCGCGCAGCGGGACTACATGGAGGCAAACTGCGAAGTCTACATCATGAGCTCCGGGGAGAAGTGGTTCACCCCTGCGTAA |
|
Protein Sequence
|
MAYANSTSTESNSSRSFRHRNANTFLTYSKCSLDPEILGLSLWSKLAPWTPAYILVAREAHQDGTWHCHALAQSVRPVTTSDPRFFDVNEYHPNIQSAKSVDRVREYILKDPLCQWEKGTFVPRKKPFVPQIGESSNTRASKDDIVRDIIQHSTNKHEYLSMLQKELPYEWATKLQYFEYSANKLFPEIAEPYTNPHPPTQPDLHCYERIEEWLNFNVYQVQPQEAGRARSLYIVGPTRTGKSTWARSLGRHNYWQNNVDWSSYDEEAVLNVIDDIPFKYCPCWKQLVGCQKNYVVNPKYGKKKKVAKRSIPAVILANEDEDWLRDMTPAQRDYMEANCEVYIMSSGEKWFTPA |
