Rice latent virus 1
Basic Information
| Genus | Mastrevirus |
|---|---|
| NCBI Assembly | GCF_004130515.1 |
| Isolate | Australia |
| Release date | 2019/2/12 |
| Submitter | Kraberger,S., Geering,A.D.W., Walters,M., Martin,D.P., Varsani,A. |
| Host | |
| Download | Genome |GFF3 |PEP |CDS |
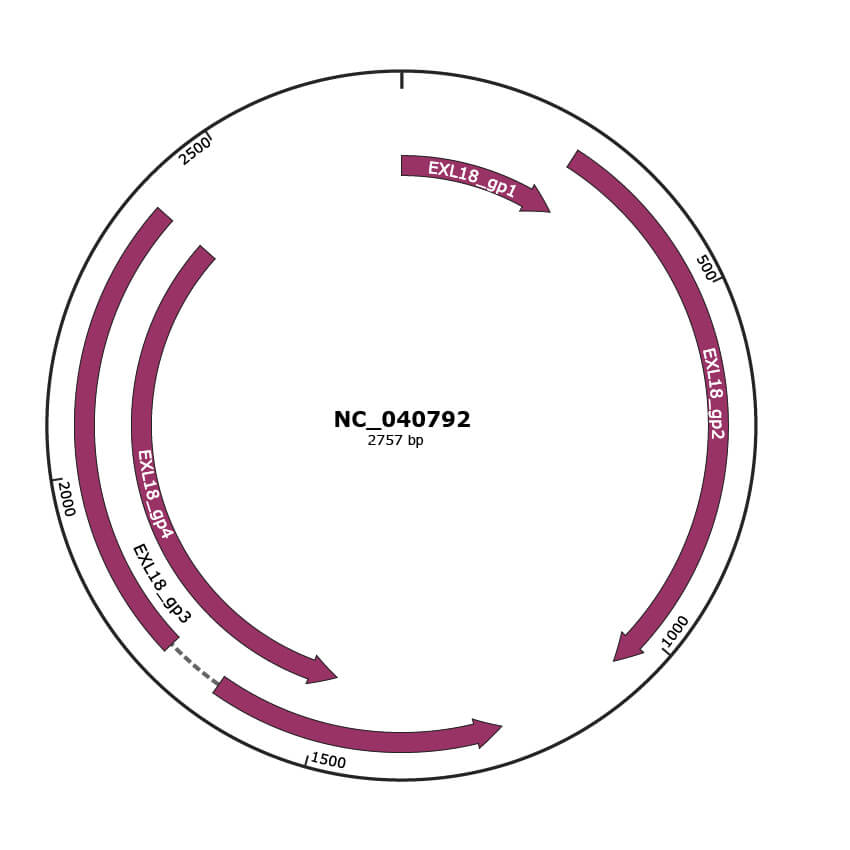
Genomic Organization
JBrowse
Genome
NC_040792
Gene Information
| NCBI Accession | YP_009553473.1 |
|---|---|
| Location | 1-267 |
| Protein Name | movement protein |
| Coding Region | ATGGCGAATTTCTCTATCCTCACTGACTCGGCCTACCCTACGGTTCCTTCGCCTCCAAGTCAAGCGGTGTCGGCCGGGTTTGAAGGTGCTCTGACGACGCTTTTCCTCGTCGTTATCGGCTCTCTACTGTTCGTTGGCGTGTTCTACCTCGGCTGGGTTCTCTGCGTGAAGGACCTCGTCCTCACCTTCAAGGCCCGCCGCCAAAGGAGGACCGAGGAGATCGGTTTCGGGAACACTCCCGCGAGGGTTGATGCCAGGGGGGTCTAA |
| Protein Sequence | MANFSILTDSAYPTVPSPPSQAVSAGFEGALTTLFLVVIGSLLFVGVFYLGWVLCVKDLVLTFKARRQRRTEEIGFGNTPARVDARGV |
| NCBI Accession | YP_009553474.1 |
|---|---|
| Location | 251-1057 |
| Protein Name | capsid protein |
| Coding Region | ATGCCAGGGGGGTCTAAGAGGGCCTCGTCCGGGAAAGATGGTTCGAGGCCGAAGCGTCGCAAGGGAGAGCCCGAGGCCGCCTTTAGGGCCCGCATATTGGGCTGGGCCTCTTCCCAGTTCGGGAGGAGGGTCTACGGTGGCGTTGGTTCGTCGCAGAGTGCTCCGCTGCAGGTGCGGACGTACAAGTGGGAGAATCCGGACAAGAGTGCCATTGCTATCGATACCGGCTCCATATGGATCGTCAATACGTTGGACGCTGGTAACGGGGATGACCAGAGACATACTCACAAGACTATTATCTACAAGATGCTGTTTCAGGGTACGATGTGGATTAATGATGCCACGTCGGCAGTCTGCGGGCCACTGACAGTTCATTTCTGGCTGGTGTATGATGCTGCGCCGAAGGGGGTTCTTCCTGCCCTTACTGACATATTCGAGGAGGGATTCTCTGGCCTGGGTTCAACCTGGGTCGTTAACCGGGACAACGTTCACCGGTTCGTGGTGAAGAGGAGGTGGCAGATGCGACTGATGACCAATGGGCAGAAGCCGTACTACACTAACCGCAGTGGGAACGGAGATCCCACCAGTGTCACGATACGGGACTTCAGGAAGTTCTTCACGAAGCTGCGTACTGCAACGGAGTGGAAGAACTCCGCAACAGGAGGGATAGGGGATATAAAGAAAGGGGCCCTATATCTTTGTGCCATGTGTTCGCAGGCACCTGAGGGGTCCGCTAGAGCTATTAACCTGGGGGTGAAGTTTAGGGGCCAGAGCCGCATATACTTCAAATGTACCGGAAATCAATGA |
| Protein Sequence | MPGGSKRASSGKDGSRPKRRKGEPEAAFRARILGWASSQFGRRVYGGVGSSQSAPLQVRTYKWENPDKSAIAIDTGSIWIVNTLDAGNGDDQRHTHKTIIYKMLFQGTMWINDATSAVCGPLTVHFWLVYDAAPKGVLPALTDIFEEGFSGLGSTWVVNRDNVHRFVVKRRWQMRLMTNGQKPYYTNRSGNGDPTSVTIRDFRKFFTKLRTATEWKNSATGGIGDIKKGALYLCAMCSQAPEGSARAINLGVKFRGQSRIYFKCTGNQ |
| NCBI Accession | YP_009553475.1 |
|---|---|
| Location | 1238-1648,1735-2388 |
| Protein Name | Rep |
| Coding Region | ATGGCCGATCCCTCCACTTCCCACCAATTCAGCGTACGGACCAAGTACTTCTTCCTCACCTATCCTAAGTGTCCTGTTGATCCTTGGGAACTCCTTCACTATCTTCTTACTCTGCTCCACAGTCGTGTACCTATCTACATACTTGTTACTCGTGAGAGTCACAAAGATGGATCCTTTCACCTACACACCCTTTTACAGCTATCTAACCGCCTGGAGACCAAAGACTCTAGGTTTTTTGACTATCAGTCTTATCATCCTAACTGTCAGCCTGCTCAGAACTGCAAGCAGGTCAGAGACTACATCACCAAGCCGGGAACCGTCAGTATTGCAGAATGGGGCGAATTCTACAACGCCAAACCGTTCAGCAAGAAGACCCTCAAGAGGAGGGCCCCTGGATCTGACACCAAGGACACCAAGATGAGGAAAATCATCTCCTCGTCCACCTCGAAAACAGAGTACCTCTCTATGGTACGCCAGACATTCCCCTTCGACTGGGCAACTAGGCTGGCCCAGTTCGAGTACAGCGCAGAGAAGCTCTTCCCCACCACCGTAACTCCATACGCAAGCTCTGTCCCCACTGATCAGCTACGTTGCAGCGAACAGCTGAGTCAGTGGGCCACAGAGAACATAAAGACATACACGGTATGTGAAGAAGATGCTCCAACAACCGCCAGAAGATATACCTCCCTATATGTATGCGGACCAACCCGAACAGGAAAGACAACCTGGGCAAGAAGCCTCGGGAAGCACAACTATTGGAACGGACTGGTGGACTTCACCACTTATGACCCCACCGCACAATACAATGTTATAGACGACATCCCGTTCAAGTACTGCCCTAATTGGAAGCAGCTAATCGGATGTCACCGTGACTACACCGTAAATCCTAAGTACGGCAAGCGGAAGGTAATAAGTGGTGGGATCCCCTCCATCATCCTGGTAAATGAAGACGAAGACTGGCTGAGTCAATGTACACCGGCTCAGACCAAGTACATCGAAAGCAACTGCGTGATACATTACATGTATGAAGGGGATAGTTTCTTCAAGCACCCTTCGGGTGCTTAG |
| Protein Sequence | MADPSTSHQFSVRTKYFFLTYPKCPVDPWELLHYLLTLLHSRVPIYILVTRESHKDGSFHLHTLLQLSNRLETKDSRFFDYQSYHPNCQPAQNCKQVRDYITKPGTVSIAEWGEFYNAKPFSKKTLKRRAPGSDTKDTKMRKIISSSTSKTEYLSMVRQTFPFDWATRLAQFEYSAEKLFPTTVTPYASSVPTDQLRCSEQLSQWATENIKTYTVCEEDAPTTARRYTSLYVCGPTRTGKTTWARSLGKHNYWNGLVDFTTYDPTAQYNVIDDIPFKYCPNWKQLIGCHRDYTVNPKYGKRKVISGGIPSIILVNEDEDWLSQCTPAQTKYIESNCVIHYMYEGDSFFKHPSGA |
| NCBI Accession | YP_009553476.1 |
|---|---|
| Location | 1489-2388 |
| Protein Name | RepA |
| Coding Region | ATGGCCGATCCCTCCACTTCCCACCAATTCAGCGTACGGACCAAGTACTTCTTCCTCACCTATCCTAAGTGTCCTGTTGATCCTTGGGAACTCCTTCACTATCTTCTTACTCTGCTCCACAGTCGTGTACCTATCTACATACTTGTTACTCGTGAGAGTCACAAAGATGGATCCTTTCACCTACACACCCTTTTACAGCTATCTAACCGCCTGGAGACCAAAGACTCTAGGTTTTTTGACTATCAGTCTTATCATCCTAACTGTCAGCCTGCTCAGAACTGCAAGCAGGTCAGAGACTACATCACCAAGCCGGGAACCGTCAGTATTGCAGAATGGGGCGAATTCTACAACGCCAAACCGTTCAGCAAGAAGACCCTCAAGAGGAGGGCCCCTGGATCTGACACCAAGGACACCAAGATGAGGAAAATCATCTCCTCGTCCACCTCGAAAACAGAGTACCTCTCTATGGTACGCCAGACATTCCCCTTCGACTGGGCAACTAGGCTGGCCCAGTTCGAGTACAGCGCAGAGAAGCTCTTCCCCACCACCGTAACTCCATACGCAAGCTCTGTCCCCACTGATCAGCTACGTTGCAGCGAACAGCTGAGTCAGTGGGCCACAGAGAACATAAAGACATACACGGTATGTGAAGAAGTATACACACAAACACGTCAAAACATGGATTATCTTGATGTAACTACTGCCGAATATCTACAGCACGTCCATGATTTGACATCCAGGATGCTCCAACAACCGCCAGAAGATATACCTCCCTATATGTATGCGGACCAACCCGAACAGGAAAGACAACCTGGGCAAGAAGCCTCGGGAAGCACAACTATTGGAACGGACTGGTGGACTTCACCACTTATGACCCCACCGCACAATACAATGTTATAG |
| Protein Sequence | MADPSTSHQFSVRTKYFFLTYPKCPVDPWELLHYLLTLLHSRVPIYILVTRESHKDGSFHLHTLLQLSNRLETKDSRFFDYQSYHPNCQPAQNCKQVRDYITKPGTVSIAEWGEFYNAKPFSKKTLKRRAPGSDTKDTKMRKIISSSTSKTEYLSMVRQTFPFDWATRLAQFEYSAEKLFPTTVTPYASSVPTDQLRCSEQLSQWATENIKTYTVCEEVYTQTRQNMDYLDVTTAEYLQHVHDLTSRMLQQPPEDIPPYMYADQPEQERQPGQEASGSTTIGTDWWTSPLMTPPHNTML |
References More References in PubMed
| 1 |
Simultaneous Detection of Herpes Simplex Virus Type 1 Latent and Lytic Transcripts in Brain Tissue. Zhang S, et al. ASN Neuro. 2022 Jan-Dec;14:17590914211053505. doi: 10.1177/17590914211053505. PMID: 35164537 |
|---|---|
| 2 |
Sanders LS 3rd, et al. J Virol. 2023 Jul 27;97(7):e0195722. doi: 10.1128/jvi.01957-22. Epub 2023 Jun 13. PMID: 37310267 |
| 3 |
Turpin JA, et al. Antimicrob Agents Chemother. 1998 Mar;42(3):487-94. doi: 10.1128/AAC.42.3.487. PMID: 9517921 |
| 4 |
Molecular characterization of a novel benyvirus infecting wheat in China. Guo J, et al. Arch Virol. 2023 Nov 6;168(12):284. doi: 10.1007/s00705-023-05912-5. PMID: 37930401 |
| 5 |
Cyclin-dependent kinases as therapeutic targets for HIV-1 infection. Rice AP. Expert Opin Ther Targets. 2016 Dec;20(12):1453-1461. doi: 10.1080/14728222.2016.1254619. Epub 2016 Nov 10. PMID: 27797603 |
| 6 |
Rice SA. Viruses. 2021 Nov 30;13(12):2395. doi: 10.3390/v13122395. PMID: 34960664 |
| 7 |
Saha JM, et al. AIDS Res Hum Retroviruses. 2018 Jan;34(1):103-110. doi: 10.1089/AID.2017.0077. Epub 2017 Dec 5. PMID: 29084447 |
| 8 |
Li C, et al. Virus Genes. 2020 Feb;56(1):67-77. doi: 10.1007/s11262-019-01708-5. Epub 2019 Oct 23. PMID: 31646461 |
| 9 |
Hajano JU, et al. Virus Res. 2015 Oct 2;208:146-55. doi: 10.1016/j.virusres.2015.06.015. Epub 2015 Jun 23. PMID: 26116274 |
| 10 |
Roles of CDKs in RNA polymerase II transcription of the HIV-1 genome. Rice AP. Transcription. 2019 Apr;10(2):111-117. doi: 10.1080/21541264.2018.1542254. Epub 2018 Nov 15. PMID: 30375919 |