Tomato leaf curl Kerala virus
Basic Information
| Genus |
Begomovirus
|
| NCBI Assembly |
GCF_000881415.1 |
| Isolate |
India |
| Release date |
2015/2/22 |
| Submitter |
Mukhopadhyay,S., Pandey,P., Nazeem,P.A., Mukherjee,S.K. |
| Host |
|
| Download |
Genome
|GFF3
|PEP
|CDS |
Genomic Organization
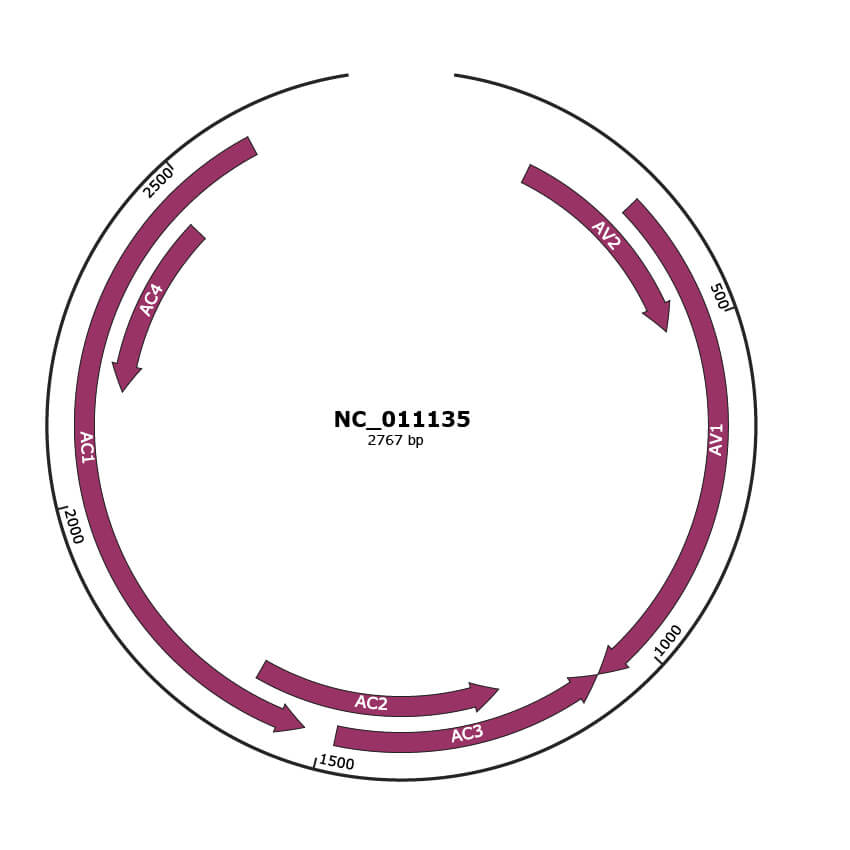
JBrowse
Genome
ACCGGATGGCCGCGAATTTTTTTGTGGGCCCCACAACGCACTAACTGACAACGACTAGTGGACCAATCACATGACGCGCTCAAAGCTTAATTGTTTTGTGGTCCCCTATTTATACTTCGGCACTAAGTATTGTACATTGCAAAATGTGGGATCCATTGTTAAACGAATTTCCCGACACTGTTCACGGTTTTAGATGTATGTTAGCAATTAAATATCTACAGTTAGTAGAAAACACTTATTCTCCTGATACATTAGGTTACGATTTAATTAGGGATTTAATTTCAGTTATTAGGGCTAGAAATTATGTCGAAGCGACCAGCAGATATAATCATTTCCACACCCGCTTCGAAGGTACGCCGCCGTCTCAACTTCGACAGCCCATATGTCAGCCGTGCTGCTGCCCCCATTGTCCGCGTCACAAAGTCAAGAGCATGGGCGAACAGGCCCATGAACAGAAAGCCCAGGATGTACAGGATGTACAGAAGCCCAGATGTCCCTAGAGGATGTGAAGGCCCATGTAAGGTACAATCATTTGAGCAAAGGCATGATATTTCTCATATTGGGAAGGTCATGTGTATTAGTGACGTTACTCGTGGGATTGGGCTAACTCATAGGGTTGGTAAGCGTTTCTGTGTTAAATCCGTTTACGTTTTGGGTAAGATATGGATGGATGAGAATATCAAGACCAAGAATCACACGAATAGTGTTATGTTTTTTCTTGTTCGTGATCGTCGTCCTGTTGACAAGCCACAAGACTTTGGAGAGGTGTTCAATATGTTTGACAATGAGCCTAGCACTGCTACTGTGAAGAATGTGCATCGAGATCGTTATCATGTTTTGAGGAAGTGGCATGCAACTGTCACTGGTGGTCAGTACGCTTCAAAGGAACAGGCATTAGTGAAGAAGTTTGTTAACGTAATAATTTATGTTGTTTATAACCAGCAAGAGGCTGGGAAATATGAGAATCATTCTGAGAATGCTTTGATGTTGTATATGGCATGTACTCATGCCTCTAATTCTGTGTACGCTACGCTGAAGATACGTATCTACTTCTATGATTCTGTAACAAACTGAATTAATAAAGATTGTATTTTATTGAATATGATTGGTCTACATATACAATGTGGTGTAATACATTCCATAATACATGATCAACAGCTCTAATTACATTGTTAATACTGATGACTCCTAAATTATTTAAATATTTATACACTTGGGTCTTAAAGACTCTTAAGAAACGACCAGTCTGAGGCTGTGAGGTCATCCAGATTCGGAAGACTAGGAAACATTTGTGTATCCCCAACACTTTCCTCAGGTTGTGATTGAACTGTACTTGGACGGTGATGATGTCTTGGTTCATCAGGAATGGCCGGTTGTGGTGTTCTGTTATCTTGAAAAATAGGGGATTTTGAATCTCCCAGATAAACACGCCATTCTCTGCTTGAGCTGCAGTGATGGGTTCCCCTGTGCGTGAATCCATGGTTGTGGCAGGCTAATGCAATGAAGTATGAACAGCCGCACGGTAGATCAACACGTCGACGCCTGGTCCCCTTCTTGGCTAGCCTGTGCTGCACTTTGATTGGTACCTGAGTACAGTGGGCCTTCGAGGGTGACGAAGATCGCATTCTTTAATGCCCAGTTTTTTAGTGCAGAATTCTTATCTTCATCCAAGAACTCTTTATAGCTGGAATTTGGTCCTGGATTGCAGAGGAAGATAGTGGGAATTCCGCCTTTAATTTGAACTGGCTTCCCGTACTTTGTATTTGATTGCCAGTCCCTTTGGGCCCCCATGAATTCTTTAAAGTGCTTTAGATAGTGGGGGTCTACGTCATCAATGACGTTATACCAGGCATCATTACTGTAGACCTTTGGACTAAGGTCTAAATGACCACACAGATAATTATGTGGTCCCAGTGATCTGGCCCACATAGTCTTTCCCGTACTACTATCACCCCTCATGACTATACTTACAGGTCTCAAAGGCCGCGCAGCGGCACTGACGACGTTCTCAGCAGCCCACTCTTCAAGTTCTTCTGGAACTTGATCAAGGAAGAAAGAAAAAAAGGGGGAAACATAACCTTCTTTAGGAAGTGTAAAATCCCTATCTAAATTAAGATCTAATAATGGAAATTGGAAAACAAAACCTTGGGGTGCTAATTCTTTAATTACTCTAAGAGCCTCTGACTTACTGCCTGCGTTAAGCGCTGCGGCGTAAGCATCATACGCTGACTGTGACCCCCCCTCTGCAGATCTCCATTCGATCTGAAACTCTCCCCAGTCGAGGGTGTCTCCGTCCTTCTCCAGATAGGACTTGACGTCTGAGCTTGATTTAGCTCCCTGAATGTTCGGATGGAAATGTGCTGACCTGGTTGGGGATACCAGGTCGAAGAATCTGTTATTTTGGCACTTGTATTTTCCTTCGAACTGGATGAGCACGTGAAGATGAGGTTCCCCATTTTCATGAAGCTCTCTGCAGATTTTTATGTATTTTTTGTTGGTTGGTGTTGCTAGGTTTTGTAATTGGGAAAGTGCTTCTTCTTTGGTGAGAGAGCATTTGGGATATGTTAGGAAATAATTTTTGGAGTTAATAAGGAAACGCTTTGGAGGCATGTTGACTAAGATAGAGGACCCGATTGACTGCTCTTGCAACTCTCCCCTGTATATCAGGTCTCAATATATAGTGAGACCCAAATGGCATAATGGTAATTAAAAAAACATTAATTTGAAATTCAAAACAAAAGGCTAAAGCGGCCATCCGTATAATATT
Gene Information
|
NCBI Accession
|
YP_002124294.1
|
|
Location
|
144-500 |
|
Gene Name
|
AV2 |
|
Protein Name
|
pre-coat protein |
|
Coding Region
|
ATGTGGGATCCATTGTTAAACGAATTTCCCGACACTGTTCACGGTTTTAGATGTATGTTAGCAATTAAATATCTACAGTTAGTAGAAAACACTTATTCTCCTGATACATTAGGTTACGATTTAATTAGGGATTTAATTTCAGTTATTAGGGCTAGAAATTATGTCGAAGCGACCAGCAGATATAATCATTTCCACACCCGCTTCGAAGGTACGCCGCCGTCTCAACTTCGACAGCCCATATGTCAGCCGTGCTGCTGCCCCCATTGTCCGCGTCACAAAGTCAAGAGCATGGGCGAACAGGCCCATGAACAGAAAGCCCAGGATGTACAGGATGTACAGAAGCCCAGATGTCCCTAG |
|
Protein Sequence
|
MWDPLLNEFPDTVHGFRCMLAIKYLQLVENTYSPDTLGYDLIRDLISVIRARNYVEATSRYNHFHTRFEGTPPSQLRQPICQPCCCPHCPRHKVKSMGEQAHEQKAQDVQDVQKPRCP |
|
NCBI Accession
|
YP_002124295.1
|
|
Location
|
304-1074 |
|
Gene Name
|
AV1 |
|
Protein Name
|
coat protein |
|
Coding Region
|
ATGTCGAAGCGACCAGCAGATATAATCATTTCCACACCCGCTTCGAAGGTACGCCGCCGTCTCAACTTCGACAGCCCATATGTCAGCCGTGCTGCTGCCCCCATTGTCCGCGTCACAAAGTCAAGAGCATGGGCGAACAGGCCCATGAACAGAAAGCCCAGGATGTACAGGATGTACAGAAGCCCAGATGTCCCTAGAGGATGTGAAGGCCCATGTAAGGTACAATCATTTGAGCAAAGGCATGATATTTCTCATATTGGGAAGGTCATGTGTATTAGTGACGTTACTCGTGGGATTGGGCTAACTCATAGGGTTGGTAAGCGTTTCTGTGTTAAATCCGTTTACGTTTTGGGTAAGATATGGATGGATGAGAATATCAAGACCAAGAATCACACGAATAGTGTTATGTTTTTTCTTGTTCGTGATCGTCGTCCTGTTGACAAGCCACAAGACTTTGGAGAGGTGTTCAATATGTTTGACAATGAGCCTAGCACTGCTACTGTGAAGAATGTGCATCGAGATCGTTATCATGTTTTGAGGAAGTGGCATGCAACTGTCACTGGTGGTCAGTACGCTTCAAAGGAACAGGCATTAGTGAAGAAGTTTGTTAACGTAATAATTTATGTTGTTTATAACCAGCAAGAGGCTGGGAAATATGAGAATCATTCTGAGAATGCTTTGATGTTGTATATGGCATGTACTCATGCCTCTAATTCTGTGTACGCTACGCTGAAGATACGTATCTACTTCTATGATTCTGTAACAAACTGA |
|
Protein Sequence
|
MSKRPADIIISTPASKVRRRLNFDSPYVSRAAAPIVRVTKSRAWANRPMNRKPRMYRMYRSPDVPRGCEGPCKVQSFEQRHDISHIGKVMCISDVTRGIGLTHRVGKRFCVKSVYVLGKIWMDENIKTKNHTNSVMFFLVRDRRPVDKPQDFGEVFNMFDNEPSTATVKNVHRDRYHVLRKWHATVTGGQYASKEQALVKKFVNVIIYVVYNQQEAGKYENHSENALMLYMACTHASNSVYATLKIRIYFYDSVTN |
|
NCBI Accession
|
YP_002124296.1
|
|
Location
|
1076-1480 |
|
Gene Name
|
AC3 |
|
Protein Name
|
AC3 |
|
Coding Region
|
ATGGATTCACGCACAGGGGAACCCATCACTGCAGCTCAAGCAGAGAATGGCGTGTTTATCTGGGAGATTCAAAATCCCCTATTTTTCAAGATAACAGAACACCACAACCGGCCATTCCTGATGAACCAAGACATCATCACCGTCCAAGTACAGTTCAATCACAACCTGAGGAAAGTGTTGGGGATACACAAATGTTTCCTAGTCTTCCGAATCTGGATGACCTCACAGCCTCAGACTGGTCGTTTCTTAAGAGTCTTTAAGACCCAAGTGTATAAATATTTAAATAATTTAGGAGTCATCAGTATTAACAATGTAATTAGAGCTGTTGATCATGTATTATGGAATGTATTACACCACATTGTATATGTAGACCAATCATATTCAATAAAATACAATCTTTATTAA |
|
Protein Sequence
|
MDSRTGEPITAAQAENGVFIWEIQNPLFFKITEHHNRPFLMNQDIITVQVQFNHNLRKVLGIHKCFLVFRIWMTSQPQTGRFLRVFKTQVYKYLNNLGVISINNVIRAVDHVLWNVLHHIVYVDQSYSIKYNLY |
|
NCBI Accession
|
YP_002124297.1
|
|
Location
|
1221-1625 |
|
Gene Name
|
AC2 |
|
Protein Name
|
AC2 |
|
Coding Region
|
ATGCGATCTTCGTCACCCTCGAAGGCCCACTGTACTCAGGTACCAATCAAAGTGCAGCACAGGCTAGCCAAGAAGGGGACCAGGCGTCGACGTGTTGATCTACCGTGCGGCTGTTCATACTTCATTGCATTAGCCTGCCACAACCATGGATTCACGCACAGGGGAACCCATCACTGCAGCTCAAGCAGAGAATGGCGTGTTTATCTGGGAGATTCAAAATCCCCTATTTTTCAAGATAACAGAACACCACAACCGGCCATTCCTGATGAACCAAGACATCATCACCGTCCAAGTACAGTTCAATCACAACCTGAGGAAAGTGTTGGGGATACACAAATGTTTCCTAGTCTTCCGAATCTGGATGACCTCACAGCCTCAGACTGGTCGTTTCTTAAGAGTCTTTAA |
|
Protein Sequence
|
MRSSSPSKAHCTQVPIKVQHRLAKKGTRRRRVDLPCGCSYFIALACHNHGFTHRGTHHCSSSREWRVYLGDSKSPIFQDNRTPQPAIPDEPRHHHRPSTVQSQPEESVGDTQMFPSLPNLDDLTASDWSFLKSL |
|
NCBI Accession
|
YP_002124298.1
|
|
Location
|
1528-2610 |
|
Gene Name
|
AC1 |
|
Protein Name
|
AC1 |
|
Coding Region
|
ATGCCTCCAAAGCGTTTCCTTATTAACTCCAAAAATTATTTCCTAACATATCCCAAATGCTCTCTCACCAAAGAAGAAGCACTTTCCCAATTACAAAACCTAGCAACACCAACCAACAAAAAATACATAAAAATCTGCAGAGAGCTTCATGAAAATGGGGAACCTCATCTTCACGTGCTCATCCAGTTCGAAGGAAAATACAAGTGCCAAAATAACAGATTCTTCGACCTGGTATCCCCAACCAGGTCAGCACATTTCCATCCGAACATTCAGGGAGCTAAATCAAGCTCAGACGTCAAGTCCTATCTGGAGAAGGACGGAGACACCCTCGACTGGGGAGAGTTTCAGATCGAATGGAGATCTGCAGAGGGGGGGTCACAGTCAGCGTATGATGCTTACGCCGCAGCGCTTAACGCAGGCAGTAAGTCAGAGGCTCTTAGAGTAATTAAAGAATTAGCACCCCAAGGTTTTGTTTTCCAATTTCCATTATTAGATCTTAATTTAGATAGGGATTTTACACTTCCTAAAGAAGGTTATGTTTCCCCCTTTTTTTCTTTCTTCCTTGATCAAGTTCCAGAAGAACTTGAAGAGTGGGCTGCTGAGAACGTCGTCAGTGCCGCTGCGCGGCCTTTGAGACCTGTAAGTATAGTCATGAGGGGTGATAGTAGTACGGGAAAGACTATGTGGGCCAGATCACTGGGACCACATAATTATCTGTGTGGTCATTTAGACCTTAGTCCAAAGGTCTACAGTAATGATGCCTGGTATAACGTCATTGATGACGTAGACCCCCACTATCTAAAGCACTTTAAAGAATTCATGGGGGCCCAAAGGGACTGGCAATCAAATACAAAGTACGGGAAGCCAGTTCAAATTAAAGGCGGAATTCCCACTATCTTCCTCTGCAATCCAGGACCAAATTCCAGCTATAAAGAGTTCTTGGATGAAGATAAGAATTCTGCACTAAAAAACTGGGCATTAAAGAATGCGATCTTCGTCACCCTCGAAGGCCCACTGTACTCAGGTACCAATCAAAGTGCAGCACAGGCTAGCCAAGAAGGGGACCAGGCGTCGACGTGTTGA |
|
Protein Sequence
|
MPPKRFLINSKNYFLTYPKCSLTKEEALSQLQNLATPTNKKYIKICRELHENGEPHLHVLIQFEGKYKCQNNRFFDLVSPTRSAHFHPNIQGAKSSSDVKSYLEKDGDTLDWGEFQIEWRSAEGGSQSAYDAYAAALNAGSKSEALRVIKELAPQGFVFQFPLLDLNLDRDFTLPKEGYVSPFFSFFLDQVPEELEEWAAENVVSAAARPLRPVSIVMRGDSSTGKTMWARSLGPHNYLCGHLDLSPKVYSNDAWYNVIDDVDPHYLKHFKEFMGAQRDWQSNTKYGKPVQIKGGIPTIFLCNPGPNSSYKEFLDEDKNSALKNWALKNAIFVTLEGPLYSGTNQSAAQASQEGDQASTC |
|
NCBI Accession
|
YP_002124299.1
|
|
Location
|
2166-2462 |
|
Gene Name
|
AC4 |
|
Protein Name
|
AC4 |
|
Coding Region
|
ATGAAAATGGGGAACCTCATCTTCACGTGCTCATCCAGTTCGAAGGAAAATACAAGTGCCAAAATAACAGATTCTTCGACCTGGTATCCCCAACCAGGTCAGCACATTTCCATCCGAACATTCAGGGAGCTAAATCAAGCTCAGACGTCAAGTCCTATCTGGAGAAGGACGGAGACACCCTCGACTGGGGAGAGTTTCAGATCGAATGGAGATCTGCAGAGGGGGGGTCACAGTCAGCGTATGATGCTTACGCCGCAGCGCTTAACGCAGGCAGTAAGTCAGAGGCTCTTAGAGTAA |
|
Protein Sequence
|
MKMGNLIFTCSSSSKENTSAKITDSSTWYPQPGQHISIRTFRELNQAQTSSPIWRRTETPSTGESFRSNGDLQRGGHSQRMMLTPQRLTQAVSQRLLE |
