Honeysuckle yellow vein virus
Basic Information
| Genus |
Begomovirus
|
| NCBI Assembly |
GCF_000859185.1 |
| Isolate |
United Kingdom:Norfolk |
| Release date |
2015/2/13 |
| Submitter |
Briddon,R.W. |
| Host |
|
| Download |
Genome
|GFF3
|PEP
|CDS |
Genomic Organization
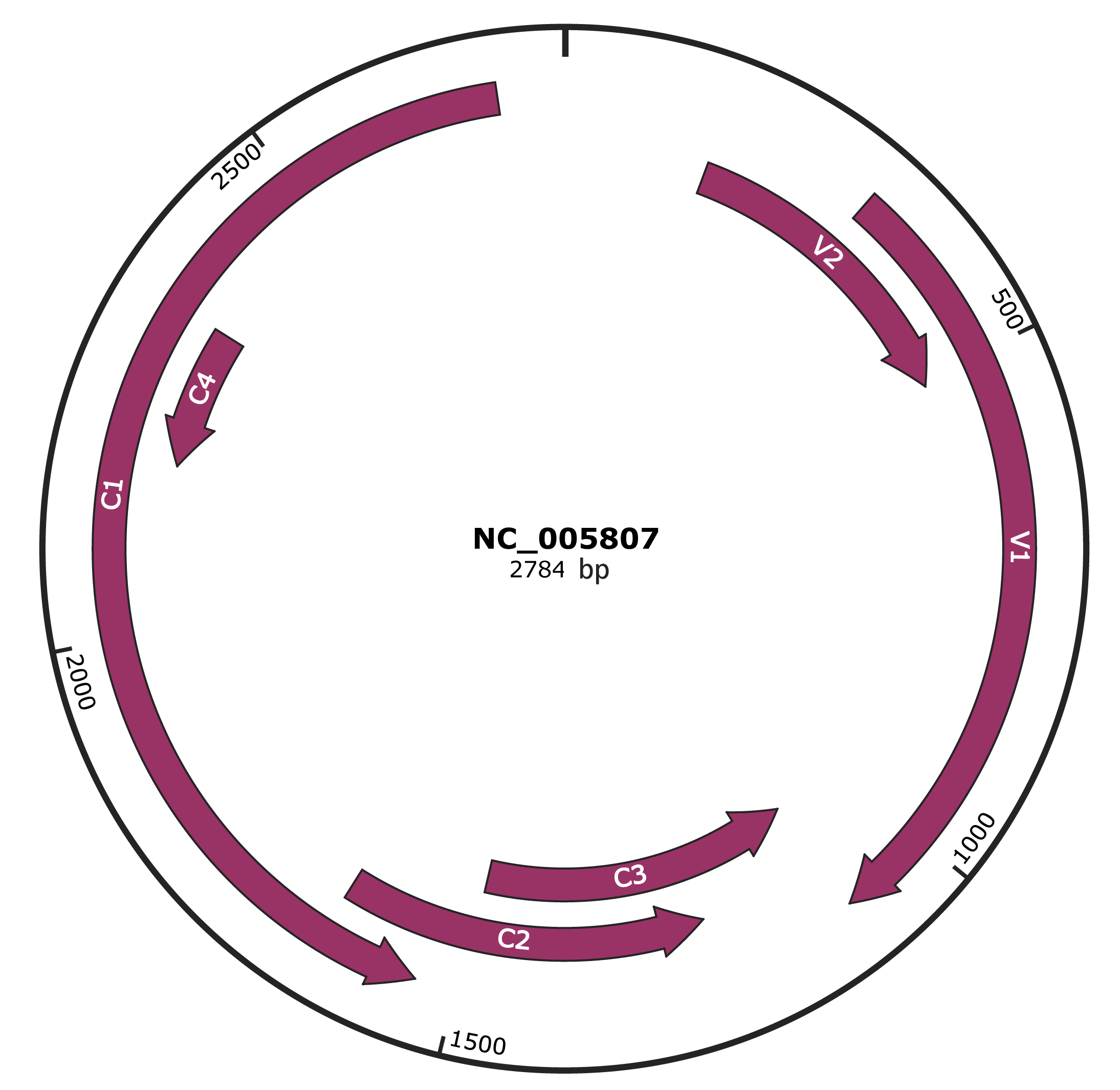
JBrowse
Genome
ACCGGATGGCCGCCCGAAATTTCGTGGTGGTCCCCCCCATGTCGGCCGTGGTCCCCCCCATGTCGGCCAATGAATAACGTGCCTTGAATGTTAGATAAGTGGAAAGTGACGTTTGCTTTTGTCTTTATATACGTCAGCCCTAATTTTAACGGTGTAAGATGTGGGATCCTTTAGTTAACGATTTCCCTGAAACCGTTCACGGTTTTAGGTGTATGTTAGCGGTTAAGTACCTCCTACTTGTTGAATCTACGTACGCTCCAGATACTGTTGGGTACGATCTTGTACGCGATCTTATTGGAGTTGTTCGAGCTAAGAACTATGTCGAAGCGTCCTGCAGATATAGGAATTTTCACACCCGTCTCCAAGGTACGGCGCCGTCTGAACTTCGACAGCCCGTATGTCAGCCGTGCGAGTGCCCCCATTGCCCTCGTCACAAACAAAAGGAGATCATGGGCTCAGCGGCCCATGTATCGGAAGCCCAGGATGTACAGAATGTACAGAAGTCCTGATGTTCCCAAGGGGTGTGAAGGTCCGTGTAAGGTACAGTCATTTGAGACGAAGAATGATGTTGGGCATTCTGGTACTCTGCTCTGTGTTTCTGATATCACCCGTGGTAATGGACTTACTCATCGTGTTGGGAAGAGATTTTGTATAAAATCTGTTTATATTATTGGTAAAATCTGGATGGATGACAATATTAAGACGAAGAATCACACTAACAACGTGTTATTCTGGTTAGTTAGGGATAGACGTCCTGGTTCAACTCCTTATGGATTCCAAGAGGCATTCAATATGATTCAAAATGAGCCCAGTACAGCCACTATTAAACAGGAGTTGAGAGATCGTTTGCAGGTGTTACATAGGTTCAGTGCGACTGTTACAGGTGGACAATATGTGTCCAAGGAACAAGCTATCATCAAGAGGTTCTGGAAGTTGAATCATCATGTCACTTACAATCATCAGGAGCAAGCTAAATATGAGAACCATACTGAAAATGCTTTGTTATTGTATATGGCGTGTACTCATGCCAGTAATCCAGTGTATGCCACATTAAAAATCAGAGTGTATTTCTACGATTCAGTACAAAATTAATAAAGCTTAAATTTTATATCTGTAAATTGTTGTACACATTCAGTTCCTTCGAATACATTATATAATACATGATCTACTGCTCTAATTACATTGTTAATTGATATTACACCCAAATTATCTAAGTACTTCAACACTTGTGTCCTAAATACCCTCAAGAAATGCAAAGTCTGAGGATGTAAGCGAGTCCAAATCTTGAAAGAGAGGAAGCACTTGTGTATGTGCAGTTGATTCCTGAGGTTGTGGTTGAATTGTATCTGTAAAGTTATTATATCCTGATCCATGTTGAGTGGCCGTTGCGCGTGTGTCGTGATTTTGAAATATAGGGGATTGGGCACTGTCCAGGTATACGCACCATTCCTTGCCTGAGCTGCAGTGATGGTGTCCCCTGTGCGTGAATCCATGATTTGCACACCCCAATGCGACGTAAATAGAACACCCGCAGTTCAGATCAATCCTCCTCCGGCGAGGTGCTCTCCTCTTCTCTGATCTGTGTCGCACCTTGATGGGTACCTGAGTAGAGTGGGCCGTAGAGGGTGACGAAGGCTGCATTTTTAATTGCCCAGGCTTTTAGGGCAGCATTCTTGTCCTCGTCTAAATACTCGGTGTATGATGACGTTGGGCCTGGATTGCAGAGGAAGATTGTTGGGATACCTCCTTTAATTTGAATTGGTTTCCCGTACTTGGTGTTGCTTTGCCAGTCCCTTTGGGCCCCCATGAACTCTTTAAAGTGCTTTAGATAGTGGGGATCCACGTCATCAATGACGTTGTACCAGGCCTCGTTGCTGTAAACCTTGGGGCTTAAGTCAAGATGGCCGCAAAGGTAGTTATGACGTGGACTCAAAGACCTGGCCCACATGGTCTTCCCTGTCCTGCTATCCCCTTCTATCACAATGCTAATGGGCCTCCATGGCCGCGCAGCGGAATCCCTGACGTTCTCGGCAGCCCATCCCTCAAGTTGCTCTGGAACTTGATCAAATGAAGAAGATAAAAAAGGAGAAACAAATTCCTCCAACGGAGGAGCAAAAATCCTATCTAAATTGCTATTTAAATTATGAAATTGTAAAACAAAATCTTTGGGAGCTTTCTCCCTTAATATACAGAGTGCCGATGACTTAGATCCTGAGTTGATTGCCTCGGCATACGCGTCGGTGGCAGACTGGCAACCTCCTCTAGCTGATCTTCCATCGACCTGGAACACTCCAAAATCAACGATGTCTCCGTCTTTCTCCATATAGGTCTTGACATCTGACGAGCTTTTAGCCGCCTGAATGTTGGGATGGAAATGTGCTGACCTGCTTGGGGATACTAGGTCGAAGAATCGTTGATTTGTGCAGTTGTATTTCCCCTCGAACTGGATGAGGACGTGGAGATGAGGGCTCCCATTTTGATGTAGCTCTCTGCAGATCCTTATGAATTTAATGTTAGTTGGTGTGTTTAATTTTTGTAATTGGGTTAGGGCTTCTTCTTTGTCGATGGAGCATTGTGGGTATGTGAGGAAATAATTTTTGGCTTTTAAAGAGAAACGCCCAGCTCTAGGCATGTTGACTTGGTCAATTGGGGGGCACTTCAAACTCCTGCTGAATGGGGGGGCATTGGGGGGCAATTTATATTCTGGGGGGCATGGCATTATTGTAATTACTTCAATTTTAAATTCAAAATTCAAATTGGTAAAGCGGCCATCCGTATAATATT
Gene Information
|
NCBI Accession
|
NP_991332.1
|
|
Location
|
159-509 |
|
Gene Name
|
V2 |
|
Protein Name
|
V2 protein |
|
Coding Region
|
ATGTGGGATCCTTTAGTTAACGATTTCCCTGAAACCGTTCACGGTTTTAGGTGTATGTTAGCGGTTAAGTACCTCCTACTTGTTGAATCTACGTACGCTCCAGATACTGTTGGGTACGATCTTGTACGCGATCTTATTGGAGTTGTTCGAGCTAAGAACTATGTCGAAGCGTCCTGCAGATATAGGAATTTTCACACCCGTCTCCAAGGTACGGCGCCGTCTGAACTTCGACAGCCCGTATGTCAGCCGTGCGAGTGCCCCCATTGCCCTCGTCACAAACAAAAGGAGATCATGGGCTCAGCGGCCCATGTATCGGAAGCCCAGGATGTACAGAATGTACAGAAGTCCTGA |
|
Protein Sequence
|
MWDPLVNDFPETVHGFRCMLAVKYLLLVESTYAPDTVGYDLVRDLIGVVRAKNYVEASCRYRNFHTRLQGTAPSELRQPVCQPCECPHCPRHKQKEIMGSAAHVSEAQDVQNVQKS |
|
NCBI Accession
|
NP_991333.1
|
|
Location
|
319-1092 |
|
Gene Name
|
V1 |
|
Protein Name
|
coat protein |
|
Coding Region
|
ATGTCGAAGCGTCCTGCAGATATAGGAATTTTCACACCCGTCTCCAAGGTACGGCGCCGTCTGAACTTCGACAGCCCGTATGTCAGCCGTGCGAGTGCCCCCATTGCCCTCGTCACAAACAAAAGGAGATCATGGGCTCAGCGGCCCATGTATCGGAAGCCCAGGATGTACAGAATGTACAGAAGTCCTGATGTTCCCAAGGGGTGTGAAGGTCCGTGTAAGGTACAGTCATTTGAGACGAAGAATGATGTTGGGCATTCTGGTACTCTGCTCTGTGTTTCTGATATCACCCGTGGTAATGGACTTACTCATCGTGTTGGGAAGAGATTTTGTATAAAATCTGTTTATATTATTGGTAAAATCTGGATGGATGACAATATTAAGACGAAGAATCACACTAACAACGTGTTATTCTGGTTAGTTAGGGATAGACGTCCTGGTTCAACTCCTTATGGATTCCAAGAGGCATTCAATATGATTCAAAATGAGCCCAGTACAGCCACTATTAAACAGGAGTTGAGAGATCGTTTGCAGGTGTTACATAGGTTCAGTGCGACTGTTACAGGTGGACAATATGTGTCCAAGGAACAAGCTATCATCAAGAGGTTCTGGAAGTTGAATCATCATGTCACTTACAATCATCAGGAGCAAGCTAAATATGAGAACCATACTGAAAATGCTTTGTTATTGTATATGGCGTGTACTCATGCCAGTAATCCAGTGTATGCCACATTAAAAATCAGAGTGTATTTCTACGATTCAGTACAAAATTAA |
|
Protein Sequence
|
MSKRPADIGIFTPVSKVRRRLNFDSPYVSRASAPIALVTNKRRSWAQRPMYRKPRMYRMYRSPDVPKGCEGPCKVQSFETKNDVGHSGTLLCVSDITRGNGLTHRVGKRFCIKSVYIIGKIWMDDNIKTKNHTNNVLFWLVRDRRPGSTPYGFQEAFNMIQNEPSTATIKQELRDRLQVLHRFSATVTGGQYVSKEQAIIKRFWKLNHHVTYNHQEQAKYENHTENALLLYMACTHASNPVYATLKIRVYFYDSVQN |
|
NCBI Accession
|
NP_991334.1
|
|
Location
|
1089-1493 |
|
Gene Name
|
C3 |
|
Protein Name
|
C3 protein |
|
Coding Region
|
ATGGATTCACGCACAGGGGACACCATCACTGCAGCTCAGGCAAGGAATGGTGCGTATACCTGGACAGTGCCCAATCCCCTATATTTCAAAATCACGACACACGCGCAACGGCCACTCAACATGGATCAGGATATAATAACTTTACAGATACAATTCAACCACAACCTCAGGAATCAACTGCACATACACAAGTGCTTCCTCTCTTTCAAGATTTGGACTCGCTTACATCCTCAGACTTTGCATTTCTTGAGGGTATTTAGGACACAAGTGTTGAAGTACTTAGATAATTTGGGTGTAATATCAATTAACAATGTAATTAGAGCAGTAGATCATGTATTATATAATGTATTCGAAGGAACTGAATGTGTACAACAATTTACAGATATAAAATTTAAGCTTTATTAA |
|
Protein Sequence
|
MDSRTGDTITAAQARNGAYTWTVPNPLYFKITTHAQRPLNMDQDIITLQIQFNHNLRNQLHIHKCFLSFKIWTRLHPQTLHFLRVFRTQVLKYLDNLGVISINNVIRAVDHVLYNVFEGTECVQQFTDIKFKLY |
|
NCBI Accession
|
NP_991335.1
|
|
Location
|
1234-1641 |
|
Gene Name
|
C2 |
|
Protein Name
|
C2 protein |
|
Coding Region
|
ATGCAGCCTTCGTCACCCTCTACGGCCCACTCTACTCAGGTACCCATCAAGGTGCGACACAGATCAGAGAAGAGGAGAGCACCTCGCCGGAGGAGGATTGATCTGAACTGCGGGTGTTCTATTTACGTCGCATTGGGGTGTGCAAATCATGGATTCACGCACAGGGGACACCATCACTGCAGCTCAGGCAAGGAATGGTGCGTATACCTGGACAGTGCCCAATCCCCTATATTTCAAAATCACGACACACGCGCAACGGCCACTCAACATGGATCAGGATATAATAACTTTACAGATACAATTCAACCACAACCTCAGGAATCAACTGCACATACACAAGTGCTTCCTCTCTTTCAAGATTTGGACTCGCTTACATCCTCAGACTTTGCATTTCTTGAGGGTATTTAG |
|
Protein Sequence
|
MQPSSPSTAHSTQVPIKVRHRSEKRRAPRRRRIDLNCGCSIYVALGCANHGFTHRGHHHCSSGKEWCVYLDSAQSPIFQNHDTRATATQHGSGYNNFTDTIQPQPQESTAHTQVLPLFQDLDSLTSSDFAFLEGI |
|
NCBI Accession
|
NP_991336.1
|
|
Location
|
1541-2719 |
|
Gene Name
|
C1 |
|
Protein Name
|
replication associated protein |
|
Coding Region
|
ATGCCATGCCCCCCAGAATATAAATTGCCCCCCAATGCCCCCCCATTCAGCAGGAGTTTGAAGTGCCCCCCAATTGACCAAGTCAACATGCCTAGAGCTGGGCGTTTCTCTTTAAAAGCCAAAAATTATTTCCTCACATACCCACAATGCTCCATCGACAAAGAAGAAGCCCTAACCCAATTACAAAAATTAAACACACCAACTAACATTAAATTCATAAGGATCTGCAGAGAGCTACATCAAAATGGGAGCCCTCATCTCCACGTCCTCATCCAGTTCGAGGGGAAATACAACTGCACAAATCAACGATTCTTCGACCTAGTATCCCCAAGCAGGTCAGCACATTTCCATCCCAACATTCAGGCGGCTAAAAGCTCGTCAGATGTCAAGACCTATATGGAGAAAGACGGAGACATCGTTGATTTTGGAGTGTTCCAGGTCGATGGAAGATCAGCTAGAGGAGGTTGCCAGTCTGCCACCGACGCGTATGCCGAGGCAATCAACTCAGGATCTAAGTCATCGGCACTCTGTATATTAAGGGAGAAAGCTCCCAAAGATTTTGTTTTACAATTTCATAATTTAAATAGCAATTTAGATAGGATTTTTGCTCCTCCGTTGGAGGAATTTGTTTCTCCTTTTTTATCTTCTTCATTTGATCAAGTTCCAGAGCAACTTGAGGGATGGGCTGCCGAGAACGTCAGGGATTCCGCTGCGCGGCCATGGAGGCCCATTAGCATTGTGATAGAAGGGGATAGCAGGACAGGGAAGACCATGTGGGCCAGGTCTTTGAGTCCACGTCATAACTACCTTTGCGGCCATCTTGACTTAAGCCCCAAGGTTTACAGCAACGAGGCCTGGTACAACGTCATTGATGACGTGGATCCCCACTATCTAAAGCACTTTAAAGAGTTCATGGGGGCCCAAAGGGACTGGCAAAGCAACACCAAGTACGGGAAACCAATTCAAATTAAAGGAGGTATCCCAACAATCTTCCTCTGCAATCCAGGCCCAACGTCATCATACACCGAGTATTTAGACGAGGACAAGAATGCTGCCCTAAAAGCCTGGGCAATTAAAAATGCAGCCTTCGTCACCCTCTACGGCCCACTCTACTCAGGTACCCATCAAGGTGCGACACAGATCAGAGAAGAGGAGAGCACCTCGCCGGAGGAGGATTGA |
|
Protein Sequence
|
MPCPPEYKLPPNAPPFSRSLKCPPIDQVNMPRAGRFSLKAKNYFLTYPQCSIDKEEALTQLQKLNTPTNIKFIRICRELHQNGSPHLHVLIQFEGKYNCTNQRFFDLVSPSRSAHFHPNIQAAKSSSDVKTYMEKDGDIVDFGVFQVDGRSARGGCQSATDAYAEAINSGSKSSALCILREKAPKDFVLQFHNLNSNLDRIFAPPLEEFVSPFLSSSFDQVPEQLEGWAAENVRDSAARPWRPISIVIEGDSRTGKTMWARSLSPRHNYLCGHLDLSPKVYSNEAWYNVIDDVDPHYLKHFKEFMGAQRDWQSNTKYGKPIQIKGGIPTIFLCNPGPTSSYTEYLDEDKNAALKAWAIKNAAFVTLYGPLYSGTHQGATQIREEESTSPEED |
|
NCBI Accession
|
NP_991337.1
|
|
Location
|
2182-2337 |
|
Gene Name
|
C4 |
|
Protein Name
|
C4 protein |
|
Coding Region
|
ATGTCAAGACCTATATGGAGAAAGACGGAGACATCGTTGATTTTGGAGTGTTCCAGGTCGATGGAAGATCAGCTAGAGGAGGTTGCCAGTCTGCCACCGACGCGTATGCCGAGGCAATCAACTCAGGATCTAAGTCATCGGCACTCTGTATATTAA |
|
Protein Sequence
|
MSRPIWRKTETSLILECSRSMEDQLEEVASLPPTRMPRQSTQDLSHRHSVY |
